Introduction
It's great to work together.... maybe you like OneZoom and want to help make it better. Maybe you've got a related idea of your own and are looking to get a head start. In any case we hope that you can benefit by reusing OneZoom code so that science, science communication and science education can all progress faster.
If you need software to visulise trees (or other hierarchies) with millions of items, there aren't many options. Mainstream tree viewing software is unlikely to work and even if it does, standard tree repsentations that feel fine for small trees will become counterintutive and hard to grasp for such large data sets. OneZoom has required a lot of novel technical and conceptual developments that we've been working on for more than a decade. You don't have to start your own visualisation project from scratch, we've made this page to share ideas about how we can work together, dispel a few misconceptions about what is possible, and hopefully lead to more collaborative efforts in future. Here are some ideas in a (non exhustive) list.
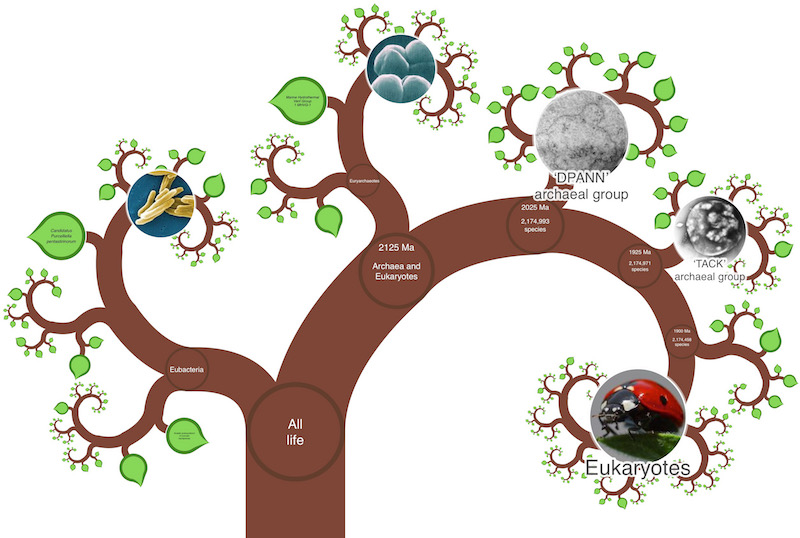
Software development
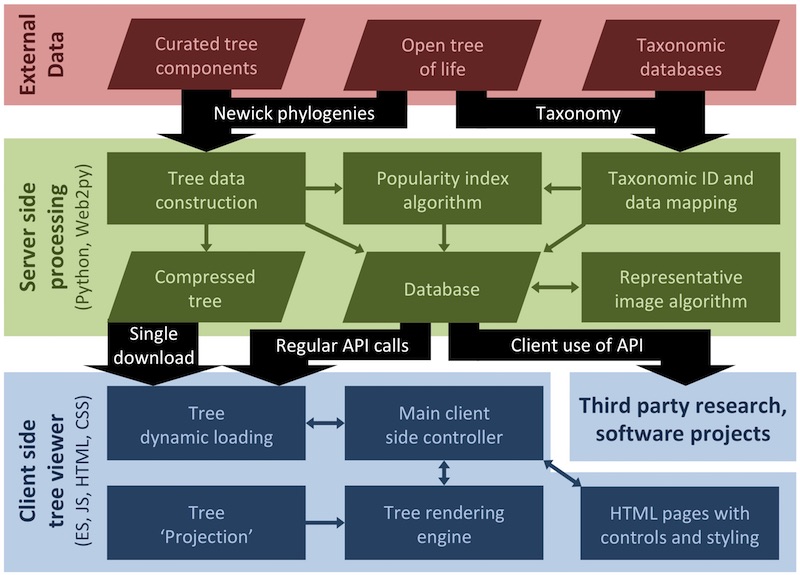
Our code is on a public GitHub repository and we provide public APIs for a range of purposes including to provide data from our index of species popularity. We also provide two Docker images of our latest codebase: oztree-complete contains everything you need to run OneZoom with no live internet connection, oztree is a download of reduced size that uses the internet to grab images as needed from the OneZoom server.
We welcome contributions from developers who are happy to help make our codebase better by contributing back some improvements. We've made a special page for developers giving a lot more detail about how to reuse or edit all or part of our codebase. A modular code structure means that different parts of the code can easily be redeveloped or swapped out, including the tree view layout, colour scheme, node and branch styles and rendering engine.
Installing a OneZoom display

We provide free display software for museums and other venues. It's easy to install because you just have to point your display machine to a web address and set up your machine so that users can go nowhere else. In case you don't have an internet connection we now provide a Docker container for our whole website, which includes the museum display page. You can find out lot more information about our museum display software on our installations page and configure a URL for your own bespoke museum display from our display launcher page. If you're planning a new exhibition and our current offering doesn't meet all your needs, you could consider budgeting for a small development project to make the necessary improvements. Please contact us if that's of interest.
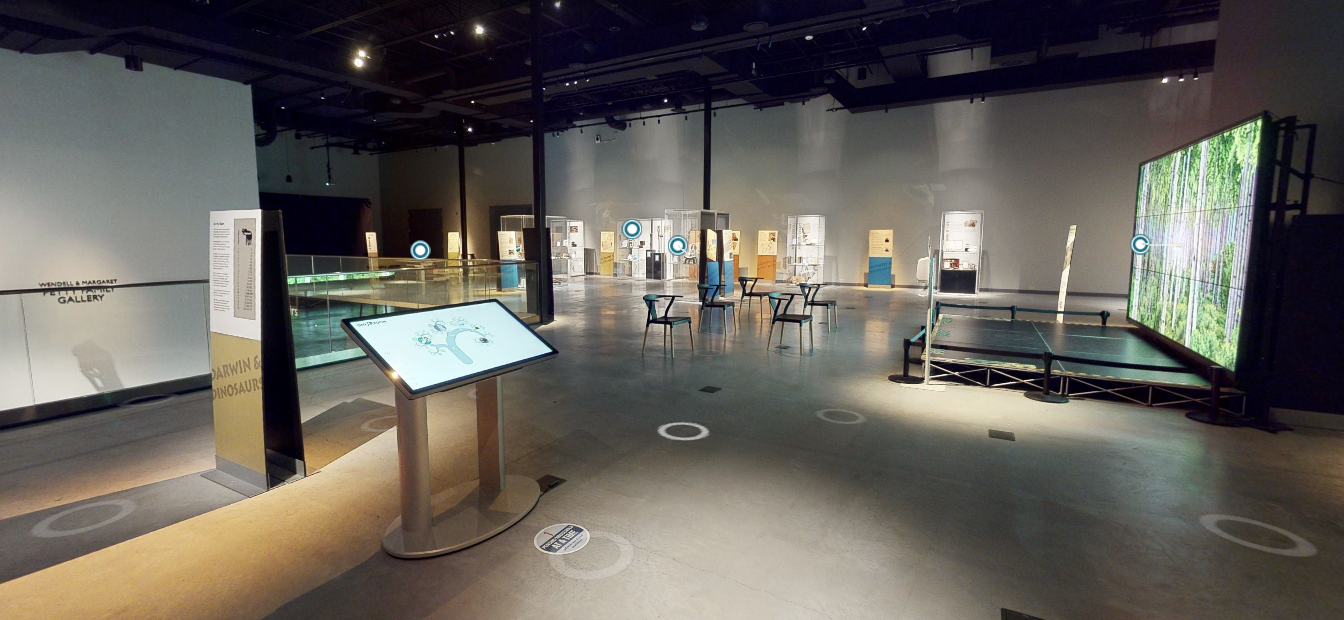

Helping with your science communication or impact
Engaging in science communication and societal impact is increasingly important for researchers, and increasingly expected by funding agencies. One way that you could do this is by collaborating with OneZoom. This doesn't necessarily mean hosting your tree using OneZoom software. For example, one project we've been planning for a while, and are actively seeking funding partners for, is an engine to take users on tours around the tree of life - a zooming animation that flies users between a number of places on the tree and gives scientific content at each stop. The content on the tour could be in the form of a narration, images or text. You could build some tours round the tree of life to communicate your research to our audience of over one million users. We're also keen to hear your own ideas.
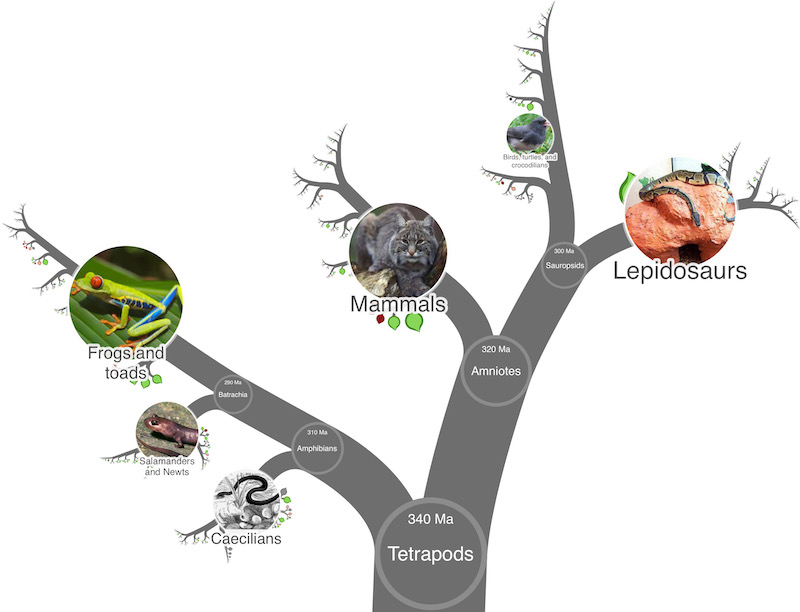
Writing OneZoom into your grant applications
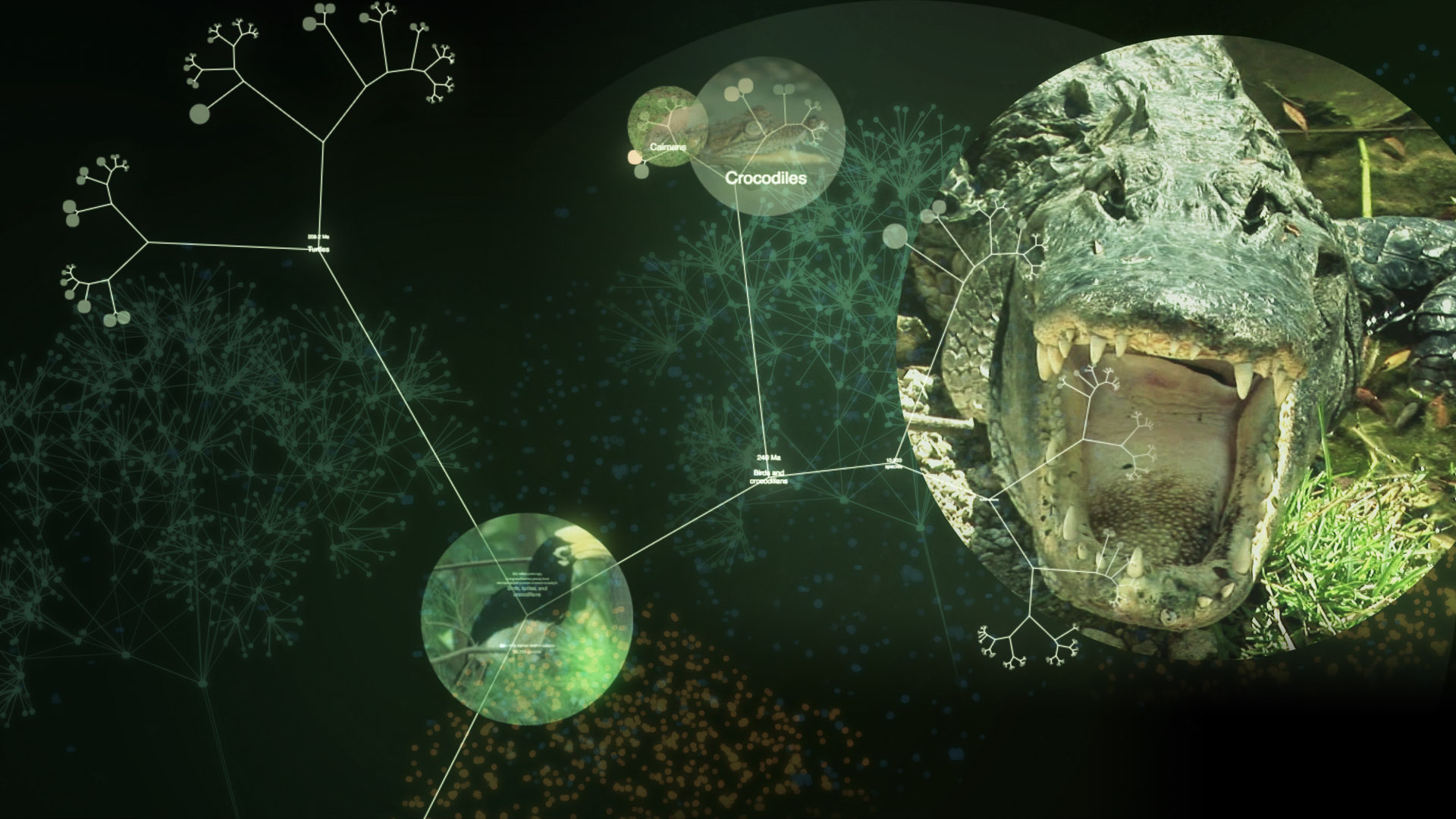
A common misconception is that OneZoom is not eligible to be included into grant applications, for example with the NSF or EU. However, because we are an independent organisation, not hosted at a university, you can include us in as a contractor in grants: this has been done already in the US and in the EU. Alternatively, you can include a third party professional developer who would do the work to improve the OneZoom codebase in the ways your project requires. One successful result of this is the One Tree One Planet project funded by the NSF. This project paid a third party developer, Jamie Lentin, to develop the OneZoom tools necessary to meet the needs of the project. Jamie has extensive experience in full stack software development, already has experience of working with the OneZoom team.

Volunteering for OneZoom
We are a charity (not-for profit) and our core team are all unpaid so we're always looking for more volunteers to help us run our website and organisation. In particular, we're looking for 'ambassadors' with experience in social media who would be happy to promote the tree of life explorer and raise awareness about its existence. We're seeking individuals with extensive experience of being a charity trustee to sit in our board meetings and share thoughts. If you're multilingual and happy to help translate the tree of life explorer into more languages that would be great. Finally, we're also looking for software developers and experienced software testers who would be happy to work on small development projects in a fork of our repository. This isn't an exhaustive list, if there's something else you'd like to do to help please let us know.
Launch your own branded tree
In you are an institution with aims aligned to our own, we can launch a branded tree for you and incorporate marked places and information on that tree specific to your organisation's aims. We do share the sponsorship revenue from such trees so if you are a not for profit organisation willing to put some work into marketing your tree it could prove another source of funding for your activities. This model has been trialled with our Linnean Society Tree
Science education
Join the growing community of teachers and educators using OneZoom in the classroom and lecture theatre to aid learning about biodiversity and evolution. If you're happy to share your educational materials and lesson plans using OneZoom please do send them to us so that we can share them with other educators. For more information see our educational materials page.
Building other tree views
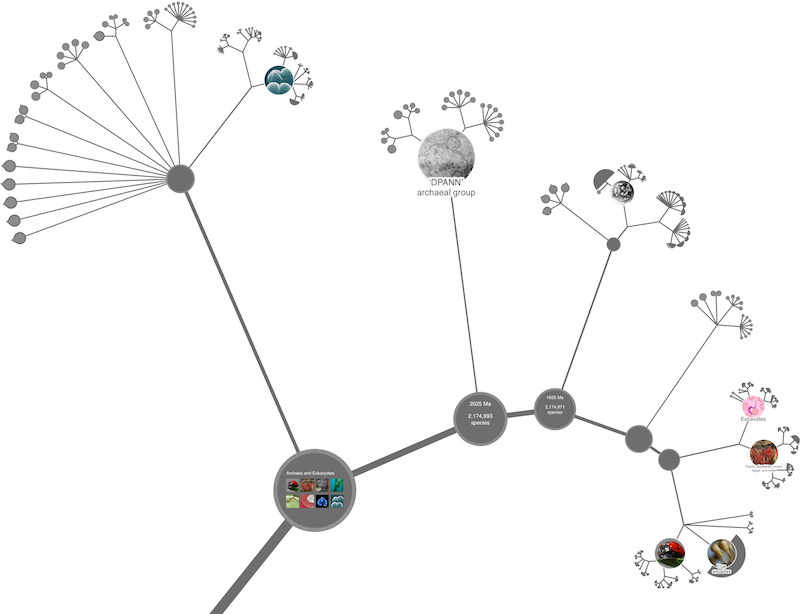
It is a common misconception that the OneZoom project is synonymous with the 'spiral' view of the tree that is our default view at present. We've dedicated a section to discussing this because we know not everyone finds the spiral view to be ideal. The OneZoom project was founded, not based on any one particular view, but on the much broader idea of fractal algorithms for laying out trees that can be explored with a zooming user interface. By a 'fractal algorithm', we mean an approach that compresses the geometry of a tree slightly as we move along branches, enabling more distant sections of the tree to be taken away from the user's focus; this generates a visualisation with a 'self similar' repeating structure. For more information you might like to read OneZoom: A Fractal Explorer for the Tree of Life. PLoS Biol 10(10): e1001406.(2012). The idea is so general that an early version of OneZoom has been forked and developed into an explorer for human genealogy available at ZoomPast.com (note that ZoomPast is an independent entity).
The part of our code that defines the points and connecting lines forming the framework of a tree visualisation is fully abstracted from the rest of the code and can be edited and replaced without worrying about any of the other modules. Do bear in mind though that to ask for a 'traditional' tree view of the complete tree of life is, in our view, a bit like asking for a traditional flat map of the whole spherical globe... it's possible to some extent, but the geometry will need adaptation and there will remain some undesirable features in one respect or another. For more information you might like Dynamic visualisation of million-tip trees: the OneZoom project (2020) bioRxiv 2020.10.14.323055
Other applications
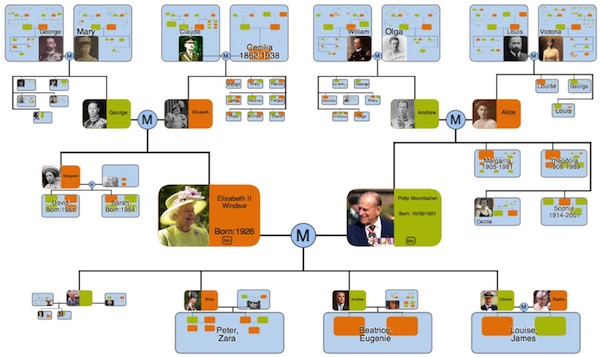
The concept of visualisation by use of a fractal inspired zoomable interface does not have to produce images that appear like a tree with leaves, and need not be restricted to the visualisation of strict hierarchies. Whilst the scope of the OneZoom organisation is currently focused on the tree of life, and adaptation to other types of data would require substantive development, below are some potential other applications of our ideas. We are interested in hearing from anyone who's actively pursuing these ideas and we might be able to offer some help.
- Monitoring of large and complex industrial processes
- Financial data, spending and performance data
- Directory structures and exploration of the contents of personal computing devices
- Complex computing structures including hardware and software
- Databases of drugs for healthcare
- Exploration of the products available for purchase from large online stores
- Family trees, genealogy and pedigrees (see ZoomPast.com)