Images
Images of species on the tree have been harvested from the internet by the Encyclopedia of Life project (EoL). We must thank EoL for their fantastic resource, and helpful responses to our various requests. We have only selected images that are in the public domain, or released under a creative commons license that allows reuse if the author is credited (and sometimes requires you to make any changes also available). To see the source for any image, zoom into the copyright symbol next to the picture. The symbol may also serve as a link to the source of the image. Exceptionally, a small number of images have been directly sent to OneZoom: copyright information is similarly accessible via the copyright symbol, and the photographer should still credited in the same manner as for any other image
IUCN Red List status
In the current version, red leaves of the tree correspond to species that are threatened with extinction according to the IUCN Red List of threatened species. Formally, these species are categorised as vulnerable, endangered, or critically endangered. Green leaves correspond to species classified as least concern, near threatened, data deficient or not evaluated. Grey leaves indicate species that are already extinct, or extinct in the wild. If you zoom in on any leaf that has been evaluated on the IUCN Red List, text should appear to indicate which category the species falls into. The categories correspond to extinction risk, which comes under our more general heading of information relevant for conservation.
Common names
Common names on the tree have been harvested from the internet by the Encyclopedia of Life project (EoL). To see the original source for a name, click on the species name in the OneZoom viewer, select the EoL tab, and from within that, click on the EoL names
tab.
Dates on the tree
Deep dates in the Archaeal stem are taken from Betts et al. (2018) Integrated genomic and fossil evidence illuminates life’s early evolution and eukaryote origin (Nature Ecology and Evolution: 2: 1556–1562. Basal eukaryotic divergences are taken from Strassert et al. (2021) A molecular timescale for eukaryote evolution with implications for the origin of red algal-derived plastids (Nature Communications: 12: 1879. Other dates are taken from The Ancestor’s Tale (2nd edition), which provides a detailed explanation in the endnotes about the dating techniques used.
Legacy trees
Additional 'legacy' trees featured on the OneZoom website may use different data and each has its own acknowledgments for data use, in the tree caption and/or embedded in the viewer itself for that particular tree.
Evolutionary Tree
The data used to construct the main tree on this site is from a hand-crafted mix of sources, but relies heavily on the Open Tree of Life project, to whom we are extremely grateful. For further, extensive information and references, please explore the tree where the data sources
menu item reveals full details about where the data came from. If you are an expert on certain species, we encourage you to contribute data to the OpenTree project.
The OneZoom tree of life is based on a bespoke backbone
, originally focussed on vertebrate species, but now extended more widely to encompass all archaea and eukaryotes.
Species have been grafted onto this backbone using a large number of studies of various groups or organisms. In particular, many studies have been collected by the Open Tree of Life project, an ambitious attempt to unify scientific studies into a comprehensive evolutionary tree of all known species of life on earth. This provides comprehensive coverage of approximately 2.2 million species, but the current draft version contains some problematic arrangements of relationships between higher-level groups (we encourage evolutionary scientists to add data to the OpenTree, from where it will eventually percolate down into the OneZoom software).
For added accuracy, and to provide a few divergence dates, the OneZoom site relies on the additional studies listed below for specific areas of the tree of life. Where possible, the OpenTree (version 13.4) has been used to supplement these studies to include most or all currently recognized species. Note that the OneZoom display currently requires all branches to split in two (bifurcations), rather than simultaneously splitting into 3 or more branches (technically known as polytomies). Where polytomies exist in the trees used, the OneZoom software splits them randomly into bifurcations.
Other sources
This page is only for information on data sources. If you're wondering how to cite OneZoom itself, or wondering about references that weren't data sources, please take a look at our About page
 Mammals: the entire tree is from Bininda-Emonds et al. (2007) The delayed rise of present-day mammals (Nature 446: 507-512), with the exception of:
Mammals: the entire tree is from Bininda-Emonds et al. (2007) The delayed rise of present-day mammals (Nature 446: 507-512), with the exception of: Marsupials: orders arranged according to Nilsson et al. (2010) Tracking Marsupial Evolution Using Archaic Genomic Retroposon Insertions (PLoS Biology: 8: e1000436).
Marsupials: orders arranged according to Nilsson et al. (2010) Tracking Marsupial Evolution Using Archaic Genomic Retroposon Insertions (PLoS Biology: 8: e1000436). Placental mammals: afrotheres and xenarthrans are grouped together (the
Placental mammals: afrotheres and xenarthrans are grouped together (the atlantogenata
hypothesis) on the basis of Poulakakis & Stamatakis (2010) Recapitulating the evolution of Afrotheria: 57 genes and rare genomic changes (RGCs) consolidate their history (Systematics and Biodiversity: 8 395-408). Cetartiodactyla: placement is taken from dos Reis et al. (2012) Phylogenomic datasets provide both precision and accuracy in estimating the timescale of placental mammal phylogeny (Proceedings of the Royal Society B: Biological Sciences 279: 3491-3500).
Cetartiodactyla: placement is taken from dos Reis et al. (2012) Phylogenomic datasets provide both precision and accuracy in estimating the timescale of placental mammal phylogeny (Proceedings of the Royal Society B: Biological Sciences 279: 3491-3500). Colugos from Janečka et al. (2008) Evidence for multiple species of Sunda colugo (Current Biology 18: R1001-R1002).
Colugos from Janečka et al. (2008) Evidence for multiple species of Sunda colugo (Current Biology 18: R1001-R1002). Primates: entire tree from Springer et al. (2012) Macroevolutionary dynamics and historical biogeography of primate diversification inferred from a species supermatrix (PLoS ONE 7: e49521 supplementary information text s2), with the exception of:
Primates: entire tree from Springer et al. (2012) Macroevolutionary dynamics and historical biogeography of primate diversification inferred from a species supermatrix (PLoS ONE 7: e49521 supplementary information text s2), with the exception of: Gibbons: arrangements of the 4 genera taken from Carbone et al. (2014) Gibbon genome and the fast karyotype evolution of small apes (Nature 513: 195-201).
Gibbons: arrangements of the 4 genera taken from Carbone et al. (2014) Gibbon genome and the fast karyotype evolution of small apes (Nature 513: 195-201).
 Sauropsids (lizards, snakes, crocodiles, birds, turtles):
Sauropsids (lizards, snakes, crocodiles, birds, turtles): Birds: entire tree from Jetz et al. (2012) The global diversity of birds in space and time (Nature 491: 444-448). See also www.birdtree.org.
Birds: entire tree from Jetz et al. (2012) The global diversity of birds in space and time (Nature 491: 444-448). See also www.birdtree.org. Snakes & lizards: arrangement and divergence dates of Gekkota, Xantusiidae, Gerrhosauridae, Cordylidae, Scincidae, Lacertoidea, Anguimorpha, Acrodonta, Pleurodonta, Serpentes, and species-level taxonomy of Sphenodon and Dibamus from Zheng & Wiens (2016) Combining phylogenomic and supermatrix approaches, and a time-calibrated phylogeny for squamate reptiles (lizards and snakes) based on 52 genes and 4162 species (Molecular Phylogenetics and Evolution 94: 537-547). Lower-level taxonomy is taken from the OpenTree.
Snakes & lizards: arrangement and divergence dates of Gekkota, Xantusiidae, Gerrhosauridae, Cordylidae, Scincidae, Lacertoidea, Anguimorpha, Acrodonta, Pleurodonta, Serpentes, and species-level taxonomy of Sphenodon and Dibamus from Zheng & Wiens (2016) Combining phylogenomic and supermatrix approaches, and a time-calibrated phylogeny for squamate reptiles (lizards and snakes) based on 52 genes and 4162 species (Molecular Phylogenetics and Evolution 94: 537-547). Lower-level taxonomy is taken from the OpenTree. Turtles: entire tree from Jaffe et al. (2011) The evolution of island gigantism and body size variation in tortoises and turtles (Biology Letters 7: 558-561)
Turtles: entire tree from Jaffe et al. (2011) The evolution of island gigantism and body size variation in tortoises and turtles (Biology Letters 7: 558-561) Crocodiles: entire tree from Oaks (2011) A time-calibrated species tree of crocodylia reveals a recent radiation of the true crocodiles (Evolution 65: 3285-97)
Crocodiles: entire tree from Oaks (2011) A time-calibrated species tree of crocodylia reveals a recent radiation of the true crocodiles (Evolution 65: 3285-97)

Lungfish: entire phylogeny and dates from Criswell (2015) The comparative osteology and phylogenetic relationships of African and South American lungfishes (Sarcoptergii:Dipnoi) (Zoological Journal of the Linnean Society 174: 801-858).
-
 Ray-finned fish: backbone from DeepFin (version 3) with the Open Tree of Life used to fill out the following orders and groups to species level: Elopiformes, Albuliformes, Notacanthiformes, Anguilliformes, Hiodontiformes, Osteoglossiformes, Clupeiformes, Alepocephaliformes, Gonorynchiformes, Cypriniformes, Gymnotiformes, Citharinoidei, Characoidei, Siluriformes, Argentiniformes, Galaxiiformes, Esociformes, Salmoniformes, Osmeriformes, Stomiiformes, Ateleopodiformes, Aulopiformes, Myctophiformes, Lampriformes, Percopsiformes, Zeiformes, Gadiformes, Polymixiiformes, Beryciformes plus Holocentriformes, Ophidiimorpharia, Batrachoidaria (Batrachoidimorpharia), Gobiomorpharia, Syngnathidae, Solenostomidae, Dactylopteroidei,Pegasidae,Callionymoidei, Pelagimorpharia, Synbranchiformes, Anabantiformes, Carangimorphariae, Ovalentaria, and Percomorpharia. For other groups, the following references have been used:
Ray-finned fish: backbone from DeepFin (version 3) with the Open Tree of Life used to fill out the following orders and groups to species level: Elopiformes, Albuliformes, Notacanthiformes, Anguilliformes, Hiodontiformes, Osteoglossiformes, Clupeiformes, Alepocephaliformes, Gonorynchiformes, Cypriniformes, Gymnotiformes, Citharinoidei, Characoidei, Siluriformes, Argentiniformes, Galaxiiformes, Esociformes, Salmoniformes, Osmeriformes, Stomiiformes, Ateleopodiformes, Aulopiformes, Myctophiformes, Lampriformes, Percopsiformes, Zeiformes, Gadiformes, Polymixiiformes, Beryciformes plus Holocentriformes, Ophidiimorpharia, Batrachoidaria (Batrachoidimorpharia), Gobiomorpharia, Syngnathidae, Solenostomidae, Dactylopteroidei,Pegasidae,Callionymoidei, Pelagimorpharia, Synbranchiformes, Anabantiformes, Carangimorphariae, Ovalentaria, and Percomorpharia. For other groups, the following references have been used: Sturgeons: entire tree from Krieger et al. (2008) The molecular phylogeny of the order Acipenseriformes revisited (Journal of Applied Ichthyology 24 Supplement s1: 36-45).
Sturgeons: entire tree from Krieger et al. (2008) The molecular phylogeny of the order Acipenseriformes revisited (Journal of Applied Ichthyology 24 Supplement s1: 36-45). Gars: all species from Deepfin 3, with Atractosteus tristoechus added from from Wright et al. (2012) Gene trees, species trees, and morphology converge on a similar phylogeny of living gars (Actinopterygii: Holostei: Lepisosteidae), an ancient clade of ray-finned fishes (Molecular Phylogenetics and Evolution 63: 848-856).
Gars: all species from Deepfin 3, with Atractosteus tristoechus added from from Wright et al. (2012) Gene trees, species trees, and morphology converge on a similar phylogeny of living gars (Actinopterygii: Holostei: Lepisosteidae), an ancient clade of ray-finned fishes (Molecular Phylogenetics and Evolution 63: 848-856). Bichirs from Suzuki et al. (2010) The mitochondrial phylogeny of an ancient lineage of ray-finned fishes (Polypteridae) with implications for the evolution of body elongation, pelvic fin loss, and craniofacial morphology in Osteichthyes (BMC Evolutionary Biology 10: 209).
Bichirs from Suzuki et al. (2010) The mitochondrial phylogeny of an ancient lineage of ray-finned fishes (Polypteridae) with implications for the evolution of body elongation, pelvic fin loss, and craniofacial morphology in Osteichthyes (BMC Evolutionary Biology 10: 209).
 Sharks, Rays, & Chimeras (cartilagenous fish): see the Chondrichthyan Tree of Life. Timing of the basal split is from Renz et al. (2013) Revealing Less Derived Nature of Cartilaginous Fish Genomes with Their Evolutionary Time Scale Inferred with Nuclear Genes (PLoS ONE 8: e66400), and species-level detail comes from the OpenTree of Life, with higher-level groupings as follows:
Sharks, Rays, & Chimeras (cartilagenous fish): see the Chondrichthyan Tree of Life. Timing of the basal split is from Renz et al. (2013) Revealing Less Derived Nature of Cartilaginous Fish Genomes with Their Evolutionary Time Scale Inferred with Nuclear Genes (PLoS ONE 8: e66400), and species-level detail comes from the OpenTree of Life, with higher-level groupings as follows: Chimaeras: arrangement of 6 major groups from Inoue et al. (2010) Evolutionary Origin and Phylogeny of the Modern Holocephalans (Chondrichthyes: Chimaeriformes): A Mitogenomic Perspective (Molecular Biology and Evolution 27: 2576-2586).
Chimaeras: arrangement of 6 major groups from Inoue et al. (2010) Evolutionary Origin and Phylogeny of the Modern Holocephalans (Chondrichthyes: Chimaeriformes): A Mitogenomic Perspective (Molecular Biology and Evolution 27: 2576-2586). Skates & Rays: arrangement of 16 higher-level groups from Aschliman et al. (2012) Body plan convergence in the evolution of skates and rays (Chondrichthyes: Batoidea) (Molecular Phylogenetics and Evolution 63: 28-42).
Skates & Rays: arrangement of 16 higher-level groups from Aschliman et al. (2012) Body plan convergence in the evolution of skates and rays (Chondrichthyes: Batoidea) (Molecular Phylogenetics and Evolution 63: 28-42). Sharks: relationships between genera (also within genera for Etmopteridae, Pristiophoridae, and Atelomycterus) primarily based on Naylor et al. (2012) Elasmobranch Phylogeny: A Mitochondrial Estimate Based on 595 Species (In: Carrier, Musick, & Heithaus (eds) Biology of Sharks and their Relatives, Edition 2. CRC Press, Boca Raton, Florida: 31-56), apart from:
Sharks: relationships between genera (also within genera for Etmopteridae, Pristiophoridae, and Atelomycterus) primarily based on Naylor et al. (2012) Elasmobranch Phylogeny: A Mitochondrial Estimate Based on 595 Species (In: Carrier, Musick, & Heithaus (eds) Biology of Sharks and their Relatives, Edition 2. CRC Press, Boca Raton, Florida: 31-56), apart from: Squaliform (deep sea) shark phylogeny from Straube et al. (2015) Molecular phylogeny of Squaliformes and first occurrence of bioluminescence in sharks (BMC Evolutionary Biology 15: 162).
Squaliform (deep sea) shark phylogeny from Straube et al. (2015) Molecular phylogeny of Squaliformes and first occurrence of bioluminescence in sharks (BMC Evolutionary Biology 15: 162).- Centroscymnus & Proscymnodon: these genera are currently excluded.

Cyclostomes
 Hagfishes: arrangement of genera from a combination of the 3 trees in Fernholm et al. (2013) Hagfish phylogeny and taxonomy, with description of the new genus Rubicundus (Craniata, Myxinidae) (Journal of Zoological Systematics and Evolutionary Research 51: 296-307).
Hagfishes: arrangement of genera from a combination of the 3 trees in Fernholm et al. (2013) Hagfish phylogeny and taxonomy, with description of the new genus Rubicundus (Craniata, Myxinidae) (Journal of Zoological Systematics and Evolutionary Research 51: 296-307). Lampreys: entire tree from figure 2.6 of Potter et al. (2015) The Taxonomy, Phylogeny, and Distribution of Lampreys (Chapter 2 in Docker (ed.) Lampreys: Biology, Conservation and Control: Volume 1. Springer, Dordrecht).
Lampreys: entire tree from figure 2.6 of Potter et al. (2015) The Taxonomy, Phylogeny, and Distribution of Lampreys (Chapter 2 in Docker (ed.) Lampreys: Biology, Conservation and Control: Volume 1. Springer, Dordrecht).
- Sea urchins, starfish, acorn worms, etc:
- Xenoturbella is a controversial group of worms, whose position is much debated. In the current version of OneZoom they are placed according to Bourlat et al. (2006) Deuterostome phylogeny reveals monophyletic chordates and the new phylum Xenoturbellida (Nature 444:85-88). Note that a more recent study places them as a basal bilaterian phylum, sister to the acoelomorph flatworms - this study also identifies five species, four of which are new to science and are shown on the OneZoom tree: see Rouse et al. (2016) New deep-sea species of Xenoturbella and the position of Xenacoelomorpha. (Nature 530: 94-97).
 The relationships between Echinozoa (sea urchins & sea cucumbers), Ophiuroidea (brittle stars), Asteroidea (starfish), and Crinoidea (sea lilies) is resolved on the basis of Pisani et al. (2012) Resolving phylogenetic signal from noise when divergence is rapid: A new look at the old problem of echinoderm class relationships (Molecular Phylogenetics and Evolution 62: 27-34).
The relationships between Echinozoa (sea urchins & sea cucumbers), Ophiuroidea (brittle stars), Asteroidea (starfish), and Crinoidea (sea lilies) is resolved on the basis of Pisani et al. (2012) Resolving phylogenetic signal from noise when divergence is rapid: A new look at the old problem of echinoderm class relationships (Molecular Phylogenetics and Evolution 62: 27-34).
 Protosomes: apart from the arrow worms, the protostomes are now commonly believed to fall into two major groups, the Ecdysozoa and the another group which sometimes gets called Lophotrochozoa or Spiralia. Within these two groups:
Protosomes: apart from the arrow worms, the protostomes are now commonly believed to fall into two major groups, the Ecdysozoa and the another group which sometimes gets called Lophotrochozoa or Spiralia. Within these two groups: Ecdysozoa (insects, and their wider relatives) now have a reasonably well established evolutionary tree.
Ecdysozoa (insects, and their wider relatives) now have a reasonably well established evolutionary tree.  The Onychophora (velvet worms) are probably closest relatives of the arthropods (crustaceans, insects, spiders, centipedes, etc), as argued by Campbell et al. (2011) MicroRNAs and phylogenomics resolve the relationships of Tardigrada and suggest that velvet worms are the sister group of Arthropoda (Proc Natl Acad Sci U S A. 108: 15920-15924). This is also suggested in the summary put together by Edgecombe et al. (2011) Higher-level metazoan relationships: recent progress and remaining questions (Organisms, Diversity, and Evolution 11:151-172). Together with the tardigrades, these form the now well-established panarthropods.
The Onychophora (velvet worms) are probably closest relatives of the arthropods (crustaceans, insects, spiders, centipedes, etc), as argued by Campbell et al. (2011) MicroRNAs and phylogenomics resolve the relationships of Tardigrada and suggest that velvet worms are the sister group of Arthropoda (Proc Natl Acad Sci U S A. 108: 15920-15924). This is also suggested in the summary put together by Edgecombe et al. (2011) Higher-level metazoan relationships: recent progress and remaining questions (Organisms, Diversity, and Evolution 11:151-172). Together with the tardigrades, these form the now well-established panarthropods. Edgecombe et al. (2011) also group the parasitic Nematomorpha (horsehair worms) with Nematoda (nematode worms). Advice from taxonomists leads us to place this as the sister to the panarthropods
Edgecombe et al. (2011) also group the parasitic Nematomorpha (horsehair worms) with Nematoda (nematode worms). Advice from taxonomists leads us to place this as the sister to the panarthropods Scalidophora (a group of tiny obscure phyla) are left as the only remaining Ecdysozoans, hence the deepest Ecdysozoan split is between them and everything else. Note that this is arguable. For instance, Scalidophora are placed as sister to nematomorpha/nematoda by Dunn et al. (2008) Broad phylogenomic sampling improves resolution of the animal tree of life Nature 452: 745-9.
Scalidophora (a group of tiny obscure phyla) are left as the only remaining Ecdysozoans, hence the deepest Ecdysozoan split is between them and everything else. Note that this is arguable. For instance, Scalidophora are placed as sister to nematomorpha/nematoda by Dunn et al. (2008) Broad phylogenomic sampling improves resolution of the animal tree of life Nature 452: 745-9.
 Spiralia (molluscs, segmented worms, and much more) — also known as Lophotrochozoa. The relationships between these animals is one of the trickiest in the entire tree. We mostly follow a recent review by Kocot (2016) On 20 years of Lophotrochozoa (Organisms Diversity & Evolution 16: 329-343), and a slighly earlier paper by Laumer et al. (2015) Spiralian Phylogeny Informs the Evolution of Microscopic Lineages (Current Biology 25: p2000-2006).
Spiralia (molluscs, segmented worms, and much more) — also known as Lophotrochozoa. The relationships between these animals is one of the trickiest in the entire tree. We mostly follow a recent review by Kocot (2016) On 20 years of Lophotrochozoa (Organisms Diversity & Evolution 16: 329-343), and a slighly earlier paper by Laumer et al. (2015) Spiralian Phylogeny Informs the Evolution of Microscopic Lineages (Current Biology 25: p2000-2006). Gnathifera, Gastrotricha and Platyhelminthes: Gnathifera is a collection of three very small, often wormy creatures, including the rotifers, which is sometimes placed with the flatworms (platyhelminths) and gastrotrichs in a group known as the
Gnathifera, Gastrotricha and Platyhelminthes: Gnathifera is a collection of three very small, often wormy creatures, including the rotifers, which is sometimes placed with the flatworms (platyhelminths) and gastrotrichs in a group known as the Platyzoa
. Although flatworms and gastrotrichs have been grouped together for almost a decade (e.g. see Dunn et al. Nature 452: 745-9), the broader idea of a coherent group of Platyzoans is increasingly thought to be wrong. We follow the alternative suggestion from Laumer et al. and Kocot that places Gnathifera as the earliest branching Spiralians, followed by the platyhelminth/gastrotrich grouping. Bryozoans (moss animals), phoronids (horseshoe worms), and brachiopods are now often found joined together in molecular studies, forming a group known as the lophophorates (see Nesnidal et al., (2013) New phylogenomic data support the monophyly of Lophophorata and an Ectoproct-Phoronid clade and indicate that Polyzoa and Kryptrochozoa are caused by systematic bias. BMC Evolutionary Biology 13: 253). Pleasingly, this is in line with older, morphological studies.
Bryozoans (moss animals), phoronids (horseshoe worms), and brachiopods are now often found joined together in molecular studies, forming a group known as the lophophorates (see Nesnidal et al., (2013) New phylogenomic data support the monophyly of Lophophorata and an Ectoproct-Phoronid clade and indicate that Polyzoa and Kryptrochozoa are caused by systematic bias. BMC Evolutionary Biology 13: 253). Pleasingly, this is in line with older, morphological studies. Entoprocta (~150 species of minute aquatic filter feeders) and Cycliophora (2 or 3 species of sac-like animals that live exclusively on lobster mouthparts) are often grouped together in molecular studies. Kocot very tentatively suggests that this group could be within the lophophorates, as sister to the bryozoans, in line with morphological studies that suggest these three phyla form a higher level group called the Polyzoa. Until a better suggestion comes along, we follow this (rather unsubstantiated) suggestion.
Entoprocta (~150 species of minute aquatic filter feeders) and Cycliophora (2 or 3 species of sac-like animals that live exclusively on lobster mouthparts) are often grouped together in molecular studies. Kocot very tentatively suggests that this group could be within the lophophorates, as sister to the bryozoans, in line with morphological studies that suggest these three phyla form a higher level group called the Polyzoa. Until a better suggestion comes along, we follow this (rather unsubstantiated) suggestion. The ribbon worms, segmented worms (annelids, e.g. earthworms) and molluscs are grouped by Edgecombe et al. with the lophophorates, in a larger band called Trochozoa. This grouping is also backed by Laumer et al. and Kocot. For the relationships between these four groups, we follow Laumer et al. in placing the basal divergence between molluscs and the rest, followed by a divergence between annelids and the remaining two. This particular arrangement is, however, extremely tentative.
The ribbon worms, segmented worms (annelids, e.g. earthworms) and molluscs are grouped by Edgecombe et al. with the lophophorates, in a larger band called Trochozoa. This grouping is also backed by Laumer et al. and Kocot. For the relationships between these four groups, we follow Laumer et al. in placing the basal divergence between molluscs and the rest, followed by a divergence between annelids and the remaining two. This particular arrangement is, however, extremely tentative. Within the segmented worms, relationships have been taken from Weigert & Bleidorn (2016) Current status of annelid phylogeny (Organisms Diversity & Evolution 16: 345-362 figure 2) rather than relying on the the OpenTree, which has particular problems with the placement of Myzostomida. Note that the OpenTree currently excludes the groups Cirratuliformia, Dinophilidae, Glyceriformia, Nerillidae, Polygordiidae, Protodrilidae, and Protodriloidae, so these are currently absent from the OneZoom tree.
Within the segmented worms, relationships have been taken from Weigert & Bleidorn (2016) Current status of annelid phylogeny (Organisms Diversity & Evolution 16: 345-362 figure 2) rather than relying on the the OpenTree, which has particular problems with the placement of Myzostomida. Note that the OpenTree currently excludes the groups Cirratuliformia, Dinophilidae, Glyceriformia, Nerillidae, Polygordiidae, Protodrilidae, and Protodriloidae, so these are currently absent from the OneZoom tree.
 Finally, the planktonic arrow worms (chaetognaths) are a perennial bugbear, but are considered by nearly all biologists to be protostomes, for example by Perez et al. (2014) The Chaetognatha: An anarchistic taxon between Protostomia and Deuterostomia (in: Wägele & Bartolomaeus (eds.) Deep Metazoan Phylogeny: The Backbone of the Tree of Life). As for the relationship between chaetognaths and the other protosomes, they have been variously placed as early-diverging ecdysozoans, spiralians, or at the base of all the protostomes — we choose the last of these after consultation with taxonomists and on the basis of a paper by Shen et al. (2015) Phylomitogenomic analyses strongly support the sister relationship of the Chaetognatha and Protostomia (Zoologica Scripta:45: 187-199).
Finally, the planktonic arrow worms (chaetognaths) are a perennial bugbear, but are considered by nearly all biologists to be protostomes, for example by Perez et al. (2014) The Chaetognatha: An anarchistic taxon between Protostomia and Deuterostomia (in: Wägele & Bartolomaeus (eds.) Deep Metazoan Phylogeny: The Backbone of the Tree of Life). As for the relationship between chaetognaths and the other protosomes, they have been variously placed as early-diverging ecdysozoans, spiralians, or at the base of all the protostomes — we choose the last of these after consultation with taxonomists and on the basis of a paper by Shen et al. (2015) Phylomitogenomic analyses strongly support the sister relationship of the Chaetognatha and Protostomia (Zoologica Scripta:45: 187-199).
 Ctenophores (comb jellies): arrangement of orders and families from Poder et al. (2001) A molecular phylogenetic framework for the phylum Ctenophora using 18S rRNA genes (Molecular Phylogenetics and Evolution 21: 218-230). On taxonomic advice, we do not follow the recent suggestion that ctenophores are the sister group to all other animals (including humans).
Ctenophores (comb jellies): arrangement of orders and families from Poder et al. (2001) A molecular phylogenetic framework for the phylum Ctenophora using 18S rRNA genes (Molecular Phylogenetics and Evolution 21: 218-230). On taxonomic advice, we do not follow the recent suggestion that ctenophores are the sister group to all other animals (including humans).- Cnidarians:
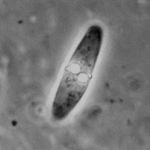 Myxozoa: placed as basal cnidarians according to Nesnidal et al. (2013) Agent of Whirling Disease Meets Orphan Worm: Phylogenomic Analyses Firmly Place Myxozoa in Cnidaria (PLoS ONE 8: e54576).
Myxozoa: placed as basal cnidarians according to Nesnidal et al. (2013) Agent of Whirling Disease Meets Orphan Worm: Phylogenomic Analyses Firmly Place Myxozoa in Cnidaria (PLoS ONE 8: e54576). Anthozoa & Medusozoa Families placed according to Collins (2009) Recent Insights into Cnidarian Phylogeny (in: Lang, M.A. et al. (2009). Proceedings of the Smithsonian Marine Science Symposium. Smithsonian Contributions to the Marine Sciences 38: 139-149).
Anthozoa & Medusozoa Families placed according to Collins (2009) Recent Insights into Cnidarian Phylogeny (in: Lang, M.A. et al. (2009). Proceedings of the Smithsonian Marine Science Symposium. Smithsonian Contributions to the Marine Sciences 38: 139-149).
 Holomycota (fungi etc.): The OpenTree has been used to fill out all the high-level fungal groups: Discicristoidea (placed at the base), Neocallimastigomycetes, Chytridiomycetes, Blastocladiomycota, Mucoromycotina, Ascomycota, Basidiomycota, Glomeromycota, Entomophthoromycota, Kickxellomycotina, Zoopagomycotina.
Holomycota (fungi etc.): The OpenTree has been used to fill out all the high-level fungal groups: Discicristoidea (placed at the base), Neocallimastigomycetes, Chytridiomycetes, Blastocladiomycota, Mucoromycotina, Ascomycota, Basidiomycota, Glomeromycota, Entomophthoromycota, Kickxellomycotina, Zoopagomycotina.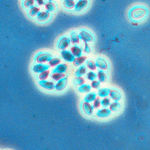 Cryptomycota and Microsporidia: these group together, according to James et al. (2013) Shared signatures of parasitism and phylogenomics unite Cryptomycota and microsporidia (Current Biology 231548-53), together with the Aphelidea (see Karpov et al. (2014) Morphology, phylogeny, and ecology of the aphelids (Aphelidea, Opisthokonta) and proposal for the new superphylum Opisthosporidia (Frontiers in Microbiology 5:112))
Cryptomycota and Microsporidia: these group together, according to James et al. (2013) Shared signatures of parasitism and phylogenomics unite Cryptomycota and microsporidia (Current Biology 231548-53), together with the Aphelidea (see Karpov et al. (2014) Morphology, phylogeny, and ecology of the aphelids (Aphelidea, Opisthokonta) and proposal for the new superphylum Opisthosporidia (Frontiers in Microbiology 5:112))-
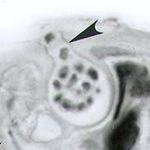 Microsporidia and allies are the deepest branching group in the fungi, followed by the Chytridiomycota, according to Capella-Gutiérrez et al. (2012) Phylogenomics supports microsporidia as the earliest diverging clade of sequenced fungi (BMC Biology 10:47).
Microsporidia and allies are the deepest branching group in the fungi, followed by the Chytridiomycota, according to Capella-Gutiérrez et al. (2012) Phylogenomics supports microsporidia as the earliest diverging clade of sequenced fungi (BMC Biology 10:47). -
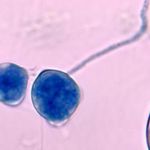 Entomophthoromycota are placed as an outgroup to Kickxellomycotina + Zoopagomycotina as a result of Gryganskyi et al. (2012) Molecular phylogeny of the Entomophthoromycota (Molecular Phylogenetics and Evolution 65:682-94).
Entomophthoromycota are placed as an outgroup to Kickxellomycotina + Zoopagomycotina as a result of Gryganskyi et al. (2012) Molecular phylogeny of the Entomophthoromycota (Molecular Phylogenetics and Evolution 65:682-94). - Other fungal groups (previously lumped into 'zygomycota' agree with the arrangement found in Porter et al (2011) Molecular phylogeny of the Blastocladiomycota (Fungi) based on nuclear ribosomal DNA (Fungal Biology 115: 381-92). Also see the summary by Stajich et al. in Current Biology 19: pR840-R845.

Eukaryotes: the position of various basal groups is often debated. This tree is based on Burki et al. (2020) The New Tree of Eukaryotes (Trends in Ecology and Evolution 35: 43-55), but with excavates grouped with Diaphoretickes into the classical
bikont
clade as in Derelle et al. (2015) Bacterial proteins pinpoint a single eukaryotic root (Proceedings of the National Academy of Sciences: 112: E693–E699, and as also shown in Strassert et al. (2021) A molecular timescale for eukaryote evolution with implications for the origin of red algal-derived plastids (Nature Communications: 12: 1879. The following orders and groups of bikonts have then been filled out to species level from the OpenTree: Excavata, Haptophyta, Centrohelida, Picozoa, Alveolata, Stramenopiles, Rhizaria, Cryptophyceae, Glaucophyta, Rhodophyceae, Chloroplastida.
Archaea are taken to contain the Eukaryotes. This is the "2 Domain" hypothesis as described in Williams & Embley (2014) Archaeal "Dark Matter" and the Origin of Eukaryotes (Genome Biology and Evolution 6: 474-481). The arrangements between higher level archaeal groups are taken from Tahon, Geesink, and Ettema (2021) Expanding Archaeal Diversity and Phylogeny: Past, Present, and Future (Annual Review of Microbiology 75:359–81), with the alteration that eukaryotes are an outgroup to the Asgardarchaeota, rather than (controversially) nested within it. Within each higher level group, archaeal species have been filled out from the OpenTree.
Further details and references
For further details and up to date academic references for OneZoom specific phylogenetic trees, please see the the OneZoom github repository, in particularOZprivate/data/OZTreeBuild/AllLife/. For references used in the Open Tree of Life, see their references page, or consult the references given for each node on their main webpage at https://tree.opentreeoflife.org/.
